Transcriptomic results have been published regarding the Shamir Medical Center longitudinal study on hyperbaric oxygen therapy (HBOT), which was conducted between 2016 and 2020 on 35 healthy adults aged 64 and older [1].
Background
Recently, we discussed misconceptions from the media on research developing on HBOT for age reversal. The researchers of this study have also recently published data to support the idea that HBOT may reverse cognitive decline.
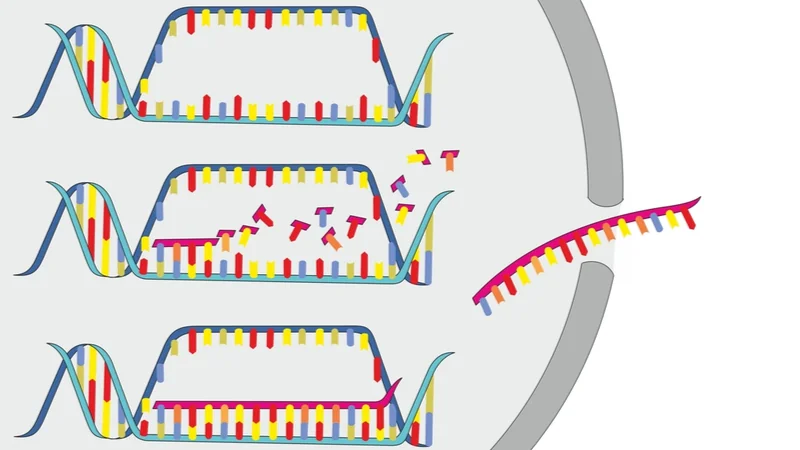
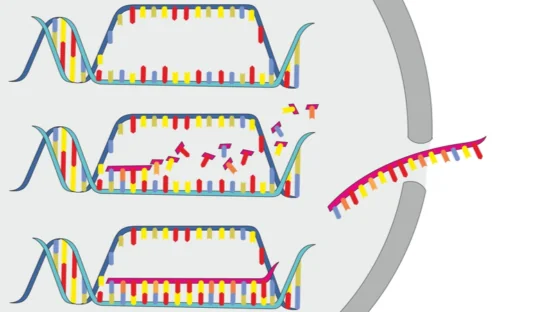
In this study, the authors sought to see if HBOT changed the transcriptome, and you can follow the link to learn more about the transcriptome.
Methods
In this study, sixty days of HBOT sessions were conducted at 100% oxygen, at 2 atmospheric pressure, for 90 minutes with five-minute air breaks every 20 minutes. The researchers examined the transcriptome using RNA extracted from whole blood samples, which were taken at a fasting baseline, at the 30th session, at the 60th session, and 1-2 weeks after the final session.
Results
Two weeks after the HBOT sessions began, compared to their baseline, 8 genes were downregulated and 11 genes were upregulated. There were nine differentially expressed genes (DEGs); however, ATP-binding cassette subfamily A member 13 (ABCA13) was the only one with over a 1.5-fold change. In studies that examine DEGs that use fold change analysis, the threshold is typically set at a 1.5-or 2-fold change [2]. The researchers verified their ABCA13 finding with the lab technique RT-qPCR on all available samples, which also yielded significant results.
The researchers continued to compare the differences from baseline at the last HBOT session and at the final extraction. The results show that there was a change in gene expression after 60 sessions of HBOT. 1342 genes were upregulated, and 570 genes were downregulated when compared to the baseline. Five of the genes had a greater than 1.5-fold change.
The authors did discuss two genes related to longevity in prior human studies [3,4,5], although their changes in expression did not meet the 1.5-fold threshold. The gene expression of FOXO3 decreased by 1.22 fold, and RUNX3, which has been shown to decline in aging [6], increased by 1.29 fold. They went on to further explain that the age-related gene expression changes of different groups of people show little overlap with each other.
Conclusion
The authors note that most of the significant changes returned to normal two weeks after the HBOT session were completed, although 19 gene expressions were still measurably different at this time. The main limitation of this study was its lack of a placebo group, so causality cannot be determined.
Due to the transcriptomic results, and the researchers’ prior results on improved cognition, it would be great to see the same research questions studied in larger sample sizes at multiple medical centers, comparing differences between people and geographical locations in order to determine if HBOT may be feasible as a therapy. We will continue to report on this as more research develops, so stay tuned.
Literature
[1] Hadanny, A., Forer, R., Volodarsky, D., Daniel-Kotovsky, M., Catalogna, M., Zemel, Y., Bechor, Y., & Efrati, S. (2021). Hyperbaric oxygen therapy induces transcriptome changes in elderly: a prospective trial. Aging, 13(undefined), 10.18632/aging.203709. Advance online publication. https://doi.org/10.18632/aging.203709
[2] Thomas, J. G., Olson, J. M., Tapscott, S. J., & Zhao, L. P. (2001). An efficient and robust statistical modeling approach to discover differentially expressed genes using genomic expression profiles. Genome research, 11(7), 1227–1236. https://doi.org/10.1101/gr.165101
[3] Flachsbart, F., Dose, J., Gentschew, L., Geismann, C., Caliebe, A., Knecht, C., Nygaard, M., Badarinarayan, N., ElSharawy, A., May, S., Luzius, A., Torres, G. G., Jentzsch, M., Forster, M., Häsler, R., Pallauf, K., Lieb, W., Derbois, C., Galan, P., Drichel, D., … Nebel, A. (2017). Identification and characterization of two functional variants in the human longevity gene FOXO3. Nature communications, 8(1), 2063. https://doi.org/10.1038/s41467-017-02183-y
[4] Flachsbart, F., Caliebe, A., Kleindorp, R., Blanché, H., von Eller-Eberstein, H., Nikolaus, S., Schreiber, S., & Nebel, A. (2009). Association of FOXO3A variation with human longevity confirmed in German centenarians. Proceedings of the National Academy of Sciences of the United States of America, 106(8), 2700–2705. https://doi.org/10.1073/pnas.0809594106
[5] Soerensen, M., Dato, S., Christensen, K., McGue, M., Stevnsner, T., Bohr, V. A., & Christiansen, L. (2010). Replication of an association of variation in the FOXO3A gene with human longevity using both case-control and longitudinal data. Aging cell, 9(6), 1010–1017. https://doi.org/10.1111/j.1474-9726.2010.00627.x
[6] Balogh, P., Adelman, E. R., Pluvinage, J. V., Capaldo, B. J., Freeman, K. C., Singh, S., Elagib, K. E., Nakamura, Y., Kurita, R., Sashida, G., Zunder, E. R., Li, H., Gru, A. A., Price, E. A., Schrier, S. L., Weissman, I. L., Figueroa, M. E., Pang, W. W., & Goldfarb, A. N. (2020). RUNX3 levels in human hematopoietic progenitors are regulated by aging and dictate erythroid-myeloid balance. Haematologica, 105(4), 905–913. https://doi.org/10.3324/haematol.2018.208918