Your #1 Source for Life Extension News
lifespan.io offers the latest information on rejuvenation biotechnology and life extension technologies. Our news outlet brings you the latest aging research, financial, and advocacy news, which is great if you like to keep up to date with everything happening in this rapidly changing field on a daily basis.
The Latest Longevity News Stories
A Popular Senolytic Treatment Causes Brain Damage in Mice
A new study calls for caution in using the well-known senolytic treatment of dasatinib and quercetin (D+Q), showing that it causes damage in certain regions of the brain, similar to what is […]

Obesity’s Effects on the Immune System May Linger for Years
A new study has suggested that T cells might retain a pro-inflammatory phenotype long after normal weight is regained following a period of obesity. In mice, […]
A Robust Senescence Response Helps Wounds Heal
A team of scientists has examined how younger and older mice heal from wounds and found that more robust senescent cell activation in younger animals helps them heal […]
Reprogrammed Cardiomyocytes Soften the Blow in Heart Attack
A new study has found that partial reprogramming mitigates the damage of myocardial infarction in mice by helping heart muscle cells to complete division [1]. When […]

It’s Springtime and the Rejuvenation Field Is Flourishing
For those of us in the Northern Hemisphere, spring is here. This is a time of renewal and hope for better times ahead, echoing […]
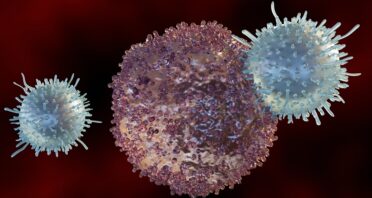
The Immune System Ages Differently in Men and Women
An investigation into the aging immune system identified age-related changes, including sex-dependent differences, in immune cell subpopulations and gene expression. In general, females showed greater age-related changes than males, including greater changes in autoimmune gene expression [1]. […]

Rapamycin Might Blunt Exercise Response in Humans
According to a new study, rapamycin probably interferes with exercise, blunting its effects in older human subjects. This result, however, might be specific to the particular protocol. Can they work together? Physical activity is one of the […]
How Inflammaging Is Linked to Epigenetic Aging
A paper in Cell Genomics has described how age-related systemic inflammation (inflammaging) is related to epigenetic aging as measured by four established clocks. Tying together two […]
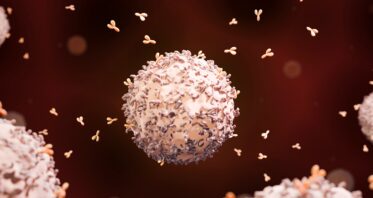
Engineered Stem Cells Become Lifelong Protein Factories
Researchers have genetically engineered blood stem cells to produce B cells that can churn out rare broad-action antibodies to fight HIV, malaria, and flu. This platform can also be used to produce other essential proteins [1]. The […]

Targeting an Appetite Hormone Receptor for Stronger Muscles
In Aging Cell, researchers have described how suppressing the ghrelin receptor improves muscle function and fights sarcopenia in older mice. An appetite hormone with […]

Vitamin C Alleviates Aging in Cynomolgus Monkeys
A recent study described a process called ferro-aging, in which iron accumulation leads to oxidative damage and cellular senescence. This process can be delayed […]

A Single Sauna Session Causes White Blood Cell Mobilization
A new study shows that hitting a sauna for 30 minutes causes a transient spike in the number of circulating white blood cells. The researchers […]
Interviewing the leading experts in aging research and longevity
We have a dedicated team of journalists who have interviewed many of the leaders in the field about their research and the drive to end age-related diseases. You can find our latest interviews below.
Regular Digest Articles
We publish the Rejuvenation Roundup – a monthly digest of what is happening in the field, a Longevity Market Recap – a monthly digest focused on the investment and business side of the field, and a quarterly Editorial – focusing on the activities of the news outlet and the wider organization.
Industry Press Releases
You can find some press releases from various companies in our field below. lifespan.io does not endorse any of these PRs and they are simply provided for information and interest.
Want even more news?
If you want to see even more recent articles, check out all news stories, or if you would like to look at a specific year or month head to the news archive.